 
Modifications to nucleotide 34 are thought to promote correct pairing to codon position 3, especially important for wobble decoding ( Agris et al. Modifications to 34 and 37 are important for balance of accuracy, flexibility, and efficiency of translation by controlling the thermodynamic and spatial limits of codon–anticodon pairing in the ribosome ( Vare et al. 2018), with wobble base 34 being most diverse. The anticodon loop (ACL) which spans positions 32–38 is most variably modified due to positions 34 and 37 ( Han and Phizicky 2018 Han et al. Several tRNAs carry t 6A37, while others are modified with either N 6-isopentenyladenosine-37 (i 6A37), a methyl addition on N 1 of G (m 1G37), or after deamination of adenosine to inosine to form m 1I37, others remain unmodified, and tRNAs Phe carry wybutosine (yW37) whose synthesis requires sequential enzymatic activities ( Lorenz et al. A relevant example is the diversity at position 37, 3′ to the anticodon. TRNA modifications comprise a wide range of chemical complexity. Because it is applicable to modifications that share a certain physiochemical characteristic, we will first provide a general review of this aspect of the method to aid understanding and its practical use. This paper reviews a method that can do so for a subset of common modifications. Sequence-specific detection of the presence or absence of modifications on specific tRNAs can be helpful in assessing the extent of suspected or actual TME dysfunction ( Abbott et al. 
Inborn errors of tRNA hypomodification lead to numerous pathological conditions of varying severity ( Chujo and Tomizawa 2021 Suzuki 2021). Some mt-tRNA modifications are not found on cy-tRNAs and vice versa ( Suzuki et al. Each of the 22-human mitochondrial (mt) tRNAs carry three to nine modifications ( Suzuki et al. Each of the ∼45 major cy-tRNA species has unique identity with a distinct combination of modified nucleotides. Human cytoplasmic (cy) tRNAs carry as many as 17 modified nucleotides, with an average of 13 per tRNA ( Pan 2018). Finally, we include a calculator tool for matching probe and hybridization conditions.Īmong all RNA species, the tRNAs are the most extensive post-transcriptionally modified both in density (% modified nucleotides) and diversity (different modifications), collectively carrying >100 modified nucleotides (nt) ( Vare et al. This is followed by data that demonstrate quantitative validation of PHAM using a TME deletion control, and that nearby modifications can falsely alter the calculated apparent modification efficiency. We provide an overview of the method and illustrate benefits of the improved conditions. Here we document these for some tRNAs and probes and illustrate principles and practices for improved reliability and uniformity in performance. Yet, over the years we have observed idiosyncratic inconsistency and variability in the assay. This positive hybridization in the absence of modification (PHAM) assay has proven useful for i 6A37, t 6A37, m 3C32, and m 2,2G26 in multiple laboratories. The signal of this probe is then normalized by a second probe elsewhere on the same tRNA. The assay method depends on differential annealing efficiency of a DNA-oligo probe to the modified versus unmodified tRNA. Our laboratory developed a northern blot method that can quantify relative levels of base modifications on multiple specific tRNAs ∼10 yr ago, which has been used to characterize four different TME deficiencies and is likely further extendable. Easily accessible methods to detect tRNA hypomodifications can facilitate progress in advancing such molecular studies. 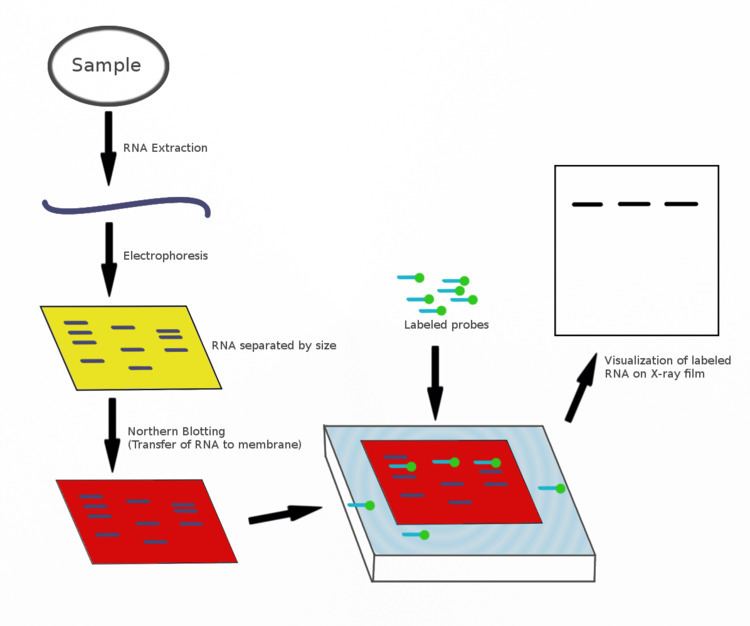
Many tRNA modifications are linked to altered gene expression, and deficiencies due to mutations in tRNA modification enzymes (TMEs) are responsible for numerous diseases. The 22 mitochondrial and ∼45 cytosolic tRNAs in human cells contain several dozen different post-transcriptional modified nucleotides such that each carries a unique constellation that complements its function.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
